ClusterType | string | | Compute cluster engine (sge or slurm). |
ClusterUser | string | | Submit username. |
ClusterQueue | string | | Queue to submit jobs. |
ClusterSubmitHost | string | | Hostname to submit jobs. |
CompleteFiles | JSON array | | JSON array of complete files, with relative paths to analysisroot. |
CreateDate | datetime | 🔴 | Date the pipeline was created. |
DataCopyMethod | string | | How the data is copied to the analysis directory: cp, softlink, hardlink. |
DependencyDirectory | string | | |
DependencyLevel | string | | |
DependencyLinkType | string | | |
Description | string | | Longer pipeline description. |
DirectoryStructure | string | | |
Directory | string | | Directory where the analyses for this pipeline will be stored. Leave blank to use the default location. |
Group | string | | ID or name of a group on which this pipeline will run |
GroupType | string | | Either subject or study |
Level | number | 🔴 | subject-level analysis (1) or group-level analysis (2). |
MaxWallTime | number | | Maximum allowed clock (wall) time in minutes for the analysis to run. |
ClusterMemory | number | | Amount of memory in GB requested for a running job. |
PipelineName | string | 🔴 🔵 | Pipeline name. |
Notes | string | | Extended notes about the pipeline |
NumberConcurrentAnalyses | number | 1 | Number of analyses allowed to run at the same time. This number if managed by NiDB and is different than grid engine queue size. |
ClusterNumberCores | number | 1 | Number of CPU cores requested for a running job. |
ParentPipelines | string | | Comma separated list of parent pipelines. |
ResultScript | string | | Executable script to be run at completion of the analysis to find and insert results back into NiDB. |
SubmitDelay | number | | Delay in hours, after the study datetime, to submit to the cluster. Allows time to upload behavioral data. |
TempDirectory | string | | The path to a temporary directory if it is used, on a compute node. |
UseProfile | bool | | true if using the profile option, false otherwise. |
UseTempDirectory | bool | | true if using a temporary directory, false otherwise. |
Version | number | 1 | Version of the pipeline. |
PrimaryScript | string | 🔴 | See details of pipeline scripts |
SecondaryScript | string | | See details of pipeline scripts. |
DataStepCount | number | 🟡 | Number of data steps. |
VirtualPath | string | 🟡 | Path of this pipeline within the squirrel package. |
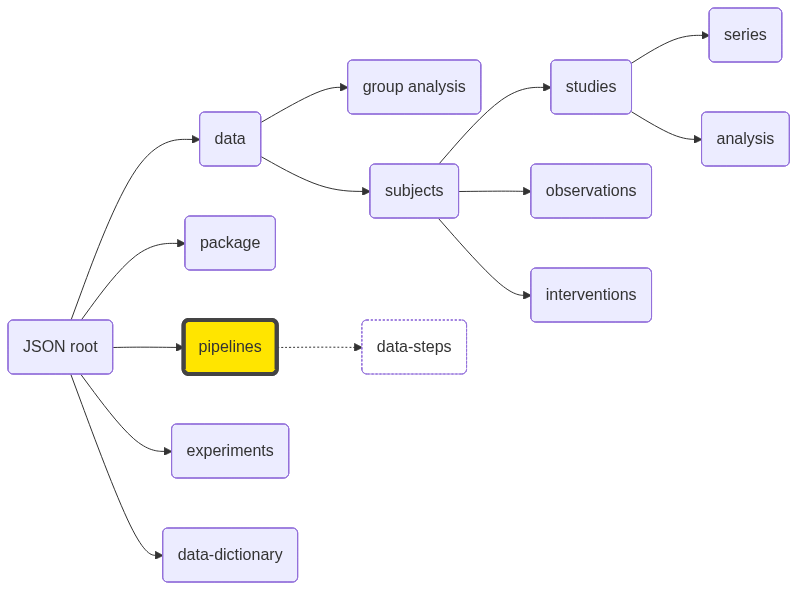
| data-steps | JSON array | | See data specifications |